1. The following
sequence is a segment from human genome. (20%)
(a) Which chromosome is this segment in?
Chromosome 18.
Perform BLAST search of this sequence in organism “Homo sapiens”, and after the
result comes out, you will see one of the entries with E value 0 is chromosome
18. So we know that this segment is in chromosome 18.
(b) What is the starting number of this segment in that chromosome? (ex.
5896432)
8935065
The first entry that BLAST returns has a LocusLink hyperlink, follow that link
and click on the LocusID entry, then find its genome annotation in the
following page, click the link to GenBank and in the accession line you can see
the start and stop position of this segment in chromosome 18.
(c) Which gene does contain this segment?
SERPINB5:
serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 5
In the LocusID entry mentioned above, you could see its name.
(d) What is the cytogenetic location of this gene? (ex. 9q34.1)
18q21.3
In the page where the LocusID entry resides, you could see its cytogenetic
location in your right hand side.
2. Use ORF finder to find the coding region of the DNA segment shown in question 1. (15%)
(a) Which frame is the correct one? What is the length of DNA and protein of this frame?
Frame “+3” is the correct one.
The length of DNA of this frame is 1128(78..1205)
The length of protein is 375aa.
(b) Give its protein sequence in fasta format.
>lcl|Sequence 1 ORF:78..1205 Frame +3
MDALQLANSAFAVDLFKQLCEKEPLGNVLFSPICLSTSLSLAQVGAKGDTANEIGQVLHFENVKDIPFGFQTVTSDVNKLSSFYSLKLIKRLYVDKSLNLSTEFISSTKRPYAKELETVDFKDKLEETKGQINNSIKDLTDGHFENILADNSVNDQTKILVVNAAYFVGKWMKKFPESETKECPFRLNKTDTKPVQMMNMEATFCMGNIDSINCKIIELPFQNKHLSMFILLPKDVEDESTGLEKIEKQLNSESLSQWTNPSTMANAKVKLSIPKFKVEKMIDPKACLENLGLKHIFSEDTSDFSGMSETKGVALSNVIHKVCLEITEDGGDSIEVPGARILQHKDELNADHPFIYIIRHNKTRNIIFFGKFCSP*
(c) Identify the protein. Give its name and accession number.
Protein name:
“serine (or cysteine) proteinase inhibitor, clade B (ovalbumin),member 5; protease inhibitor 5 (maspin) [Homo sapiens]”
Accession number: NP_0026303. Find its counterpart in mouse (M.musculus). (20%)
(a) Give its name and accession number (for protein).
Name:
serine (or cysteine) proteinase inhibitor, clade B, member 5; serine protease inhibitor 7; serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 5 [Mus musculus]
Accession number: NP_033283(NCBI)
or
Name: Maspin precursor (Protease inhibitor 5).
Accession number: P70124(Swissprot)
(b) Give the MGI number and chromosome number of its gene.
MGI: 109579
Chromosome 1.
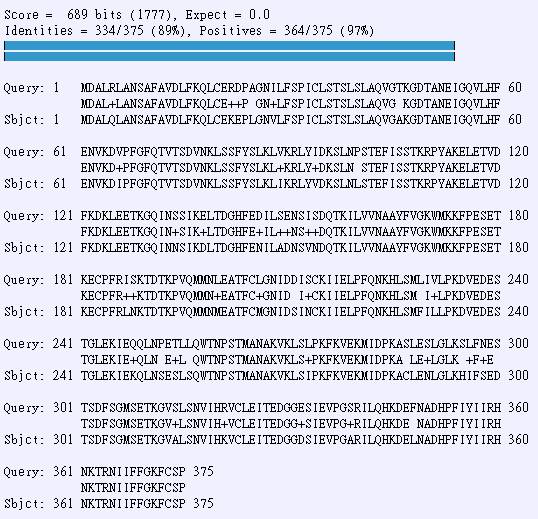
(c) Use blast2seq to compare these two protein sequences. Give % of identity and positive (Do not use filter for LCR). Show Blast2 results.
Identity: 89%(334/375)
Positives: 97%(364/375)
4. Search the conserved domains. (15%)
(a) Identify this protein in Swissprot database. Give its name and accession
number.
Name: Adenovirus 5 E1A-binding protein or BS69 protein
Accession number: Q15326
(b) Search the conserved domains that this protein may have. Give the
description and pfam or smart number of the conserved domains you found.
smart number: SM00297
description: (no description, here is the interpro abstract)
Bromodomains are found in a variety of mammalian,
invertebrate and yeast
DNA-binding proteins. Bromodomains can interact with acetylated lysine. In some
proteins, the classical bromodomain has diverged to such an extent that parts
of the region are either missing or contain an insertion (e.g., mammalian
protein HRX, C.elegans
hypothetical protein ZK783.4, yeast
protein YTA7). The bromodomain may occur as a single copy, or in duplicate.
The precise function of the domain is unclear, but it may be involved in
protein-protein interactions and may play a role in assembly or activity of
multi-component complexes involved in transcriptional activation.
smart number: SM00293
description: conservation of Pro-Trp-Trp-Pro residues
5. Analysis the above protein in ExPASy. (25%)
(a) Calculate its molecular weight and pI value.
pI=8.55
Mw=66203.27
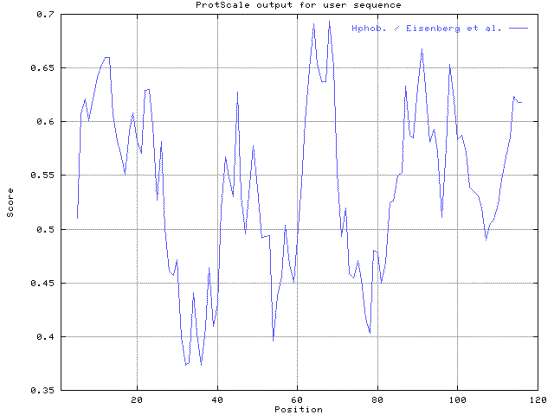
(b) Predict the hydrophobic profile of one of the domain (from 412 to 532). Use Eisenberg et al. scale, window size 9, Relative weight 90% and normalize the scale from 0 to 1. Paste the predict result on the answer sheet.
(c) Perform the trypsin cleavage for this protein. How many peptide fragments you got? Display peptides with a mass bigger than 1500 Dalton.
(d) Perform the Topology prediction. Predict this protein in SOSUI system. Give the results.
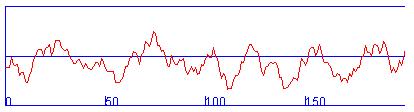
Average of hydrophobicity: -0.958541
This amino acid sequence is of a SOLUBLE PROTEIN.
Hydropathy profile:
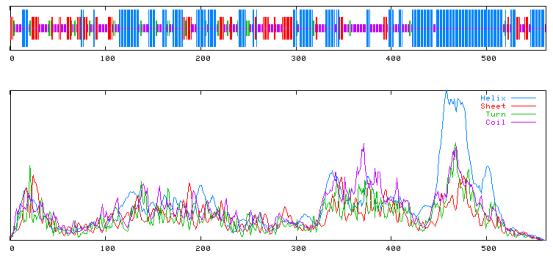
(e) Using SOPM or SOPMA to predict its secondary structure.
SOPM results:
MSRVHGMHPKETTRQLSLAVKDGLIVETLTVGCKGSKAGIEQEGYWLPGDEIDWETENHDWYCFECHLPG
heeettccccccchhhhhhhctteeeeeeeeccccccccccttteecccccceecccccceeeeetcctt
EVLICDLCFRVYHSKCLSDEFRLRDSSSPWQCPVCRSIKKKNTNKQEMGTYLRFIVSRMKERAIDLNKKGceeeehhheeeecttccchheeecccccccccceeeeecttcccchhhhhhhhhhhhhhhhhhhhhcttc
KDNKHPMYRRLVHSAVDVPTIQEKVNEGKYRSYEEFKADAQLLLHNTVIFYGADSEQADIARMLYKDTCHcccccchhhhhhhttccccchhhhhcttccchhhhhhhhhhheetteeeeeccccchhhhhhhhhhtchh
ELDELQLCKNCFYLSNARPDNWFCYPCIPNHELVWAKMKGFGFWPAKVMQKEDNQVDVRFFGHHHQRAWIhhhhhhhctteeeeecccttceeecccccthhhhhhhhttttccchheehcccccceeeeeeccccceec
PSENIQDITVNIHRLHVKRSMGWKKACDELELHQRFLREGRFWKSKNEDRGEEEAESSISSTSNEQLKVTcccccceeeeeeeeeeehhhhtcchhhhhhhhhhhhhhttceecccccccchhhhhhhhhhcchhheeee
QEPRAKKGRRNQSVEPKKEEPEPETEAVSSSQEIPTMPQPIEKVSVSTQTKKLSASSPRMLHRSTQTTNDccccccttcccccccccccccccchhhhcccccccccccccceeeechhhhhhccccchhhhccccccct
GVCQSMCHDKYTKIFNDFKDRMKSDHKRETERVVREALEKLRSEMEEEKRQAVNKAVANMQGEMDRKCKQtchhhhhhhhhhhhhhhhhhhhhhccchhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhh
VKEKCKEEFVEEIKKLATQHKQLISQTKKKQWCYNCEEEAMYHCCWNTSYCSIKCQQEHWHAEHKRTCRRhhhhhhhhhhhhhhhhhhhhhhhhhhhccthceechhhhhheeeeccccceeeecchhhhhhhhhhhhhh
KRhc
SOPM : Alpha helix (Hh) : 246 is 43.77% 310 helix (Gg) : 0 is 0.00% Pi helix (Ii) : 0 is 0.00% Beta bridge (Bb) : 0 is 0.00% Extended strand (Ee) : 92 is 16.37% Beta turn (Tt) : 40 is 7.12% Bend region (Ss) : 0 is 0.00% Random coil (Cc) : 184 is 32.74% Ambigous states (?) : 0 is 0.00% Other states : 0 is 0.00%
SOPMA results:
MSRVHGMHPKETTRQLSLAVKDGLIVETLTVGCKGSKAGIEQEGYWLPGDEIDWETENHDWYCFECHLPG
heeettccccccchhhhhhhctteeeeeeeeccccccccccttteecccccceecccccceeeeetcctt
EVLICDLCFRVYHSKCLSDEFRLRDSSSPWQCPVCRSIKKKNTNKQEMGTYLRFIVSRMKERAIDLNKKGceeeehhheeeecttccchheeecccccccccceeeeecttcccchhhhhhhhhhhhhhhhhhhhhcttc
KDNKHPMYRRLVHSAVDVPTIQEKVNEGKYRSYEEFKADAQLLLHNTVIFYGADSEQADIARMLYKDTCHcccccchhhhhhhttccccchhhhhcttccchhhhhhhhhhheetteeeeeccccchhhhhhhhhhtchh
ELDELQLCKNCFYLSNARPDNWFCYPCIPNHELVWAKMKGFGFWPAKVMQKEDNQVDVRFFGHHHQRAWIhhhhhhhctteeeeecccttceeecccccthhhhhhhhttttccchheehcccccceeeeeeccccceec
PSENIQDITVNIHRLHVKRSMGWKKACDELELHQRFLREGRFWKSKNEDRGEEEAESSISSTSNEQLKVTcccccceeeeeeeeeeehhhhtcchhhhhhhhhhhhhhttceecccccccchhhhhhhhhhcchhheeee
QEPRAKKGRRNQSVEPKKEEPEPETEAVSSSQEIPTMPQPIEKVSVSTQTKKLSASSPRMLHRSTQTTNDccccccttcccccccccccccccchhhhcccccccccccccceeeechhhhhhccccchhhhccccccct
GVCQSMCHDKYTKIFNDFKDRMKSDHKRETERVVREALEKLRSEMEEEKRQAVNKAVANMQGEMDRKCKQtchhhhhhhhhhhhhhhhhhhhhhccchhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhhh
VKEKCKEEFVEEIKKLATQHKQLISQTKKKQWCYNCEEEAMYHCCWNTSYCSIKCQQEHWHAEHKRTCRRhhhhhhhhhhhhhhhhhhhhhhhhhhhccthceechhhhhheeeeccccceeeecchhhhhhhhhhhhhh
KRhc
SOPMA :
Alpha helix (Hh) : 246 is 43.77% 310 helix (Gg) : 0 is 0.00% Pi helix (Ii) : 0 is 0.00% Beta bridge (Bb) : 0 is 0.00% Extended strand (Ee) : 92 is 16.37% Beta turn (Tt) : 40 is 7.12% Bend region (Ss) : 0 is 0.00% Random coil (Cc) : 184 is 32.74% Ambigous states (?) : 0 is 0.00% Other states : 0 is 0.00%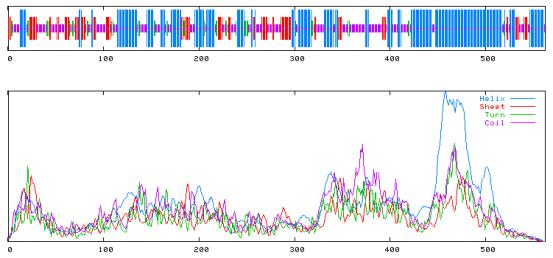
※ Bonus:
Please explain these following terms.
(1) Filter (Low-complexity) (hint: when using blast)
進行BLAST序列比對時將序列中組成成分是low complexity的區域mask掉的遮罩
(2) Matrix
進行protein-protein
BLAST序列比對時,用來計算alignment分數的給分矩陣
(3) Homologous gene
不同物種間序列相似的同源基因
(4) Contig
在shotgun
method中,數組由gel
electrophoresis得到並且互相之間有overlapping的序列為一個contig,每一組由gel electrophoresis得到的序列至少為一contig。